Bio/statistics handout 13: Eigenvalues in biology
a) An
example from genetics:
Suppose
we have a fixed size N population of cells that reproduce by fission. The model here is that there are N cells that
divide at the start, thus producing 2N cells. After one unit of time, half of these die and
half survive, so there are N cells to fission at the end of 1 unit of
time. These N survivors divide to
produce 2N cells and so start the next run of the cycle. In particular, there are always N surviving
cells at the end of one unit of time and then 2N just at the start of the next
as each of these N cells splits in half.
Now,
suppose that at the beginning, t = 0, there is, for each n Î {0,
1, …, N}, a given probability which I’ll call pn(0) for n of the N initial
cells to carry a certain trait. Suppose
that this trait (red color as opposed to blue) is neutral with respect to the
cell’s survival. In other words, the
probability is ![]() for any given red cell
to survive to reproduce, and the probability is
for any given red cell
to survive to reproduce, and the probability is ![]() for any given blue
cell to survive to reproduce.
for any given blue
cell to survive to reproduce.
Here is
the key question: Given the initial
probabilities, p0(0), p1(0), …, pN(0), what
are the corresponding probabilities after some t generations? Thus, what are the values of p0(t),
p1(t), … , pN(t) where now pn(t) denotes the
probability that n cells of generation t carry the red color.
To solve
this, we can use our tried and true recourse to conditional probabilities by
noting that
pt(n) = Prob(n red
survivors|0 red parents)·p0(t-1)
+ Prob(n red survivors|1 red parent) p1(t-1) +
Prob(n red survivors|2 red parents) p2(t-1) + ···
(13.1)
where Prob(n red survivorst|m red
parents) is the conditional probability that there are n red cells at the end
of a cycle given that there were m such cells the end of the previous
cycle. If the ambient environmental
conditions don’t change, one would explect that these conditional probabilities
are independent of time. As I explain
below, they are, in fact, computable from what we are given about this
problem. In any event, let me use the
shorthand A to denote the square matrix of size N+1 whose entry in row n and
column m is Prob(n red survivors|m red parents). In this regard, note that n and m run from 0
to N, not the from 1 to N+1. Let ![]() (t) denote the vector in RN+1 whose n’th entry
is pn(t). Then (13.1) reads
(t) denote the vector in RN+1 whose n’th entry
is pn(t). Then (13.1) reads
![]() (t) = A
(t) = A![]() (t-1) .
(t-1) .
(13.2)
This
last equation would be easy to solve if we knew that ![]() (0) was an eigenvector
of the matrix A. That is, if it were the
case that
(0) was an eigenvector
of the matrix A. That is, if it were the
case that
A![]() (0) = l
(0) = l![]() (0) with l some real number.
(0) with l some real number.
(13.3)
Indeed, were this the case, then
the t = 1 version of (13.2) would read ![]() (1) = l
(1) = l![]() (0). We could then use
the t = 2 version of (13.2) to compute
(0). We could then use
the t = 2 version of (13.2) to compute ![]() (2) and we would find that
(2) and we would find that ![]() (2) = lA
(2) = lA![]() (0) = l2
(0) = l2![]() (0). Continuing in the
vein, we would find that
(0). Continuing in the
vein, we would find that ![]() (t) = lr
(t) = lr![]() (0) and our problem would be solved.
(0) and our problem would be solved.
Now,
it is unlikely that ![]() (0) is going to be an eigenvector. However, even if
(0) is going to be an eigenvector. However, even if ![]() (0) is a linear combination of eigenvectors,
(0) is a linear combination of eigenvectors,
![]() (0) = c1
(0) = c1![]() 1 + c2
1 + c2![]() 2 + ··· ,
2 + ··· ,
(13.4)
we could still solve for ![]() (t). Indeed, let lk denote the eigenvalue for any given
(t). Indeed, let lk denote the eigenvalue for any given ![]() k. Thus, A
k. Thus, A![]() k = lk
k = lk![]() k. Granted
we know these eigenvalues and eigenvectors, then we can plug (13.4) into the t
= 1 version of (13.2) to find
k. Granted
we know these eigenvalues and eigenvectors, then we can plug (13.4) into the t
= 1 version of (13.2) to find
![]() (1) = c1l1
(1) = c1l1![]() 1 + c2l2
1 + c2l2![]() 2 + ··· .
2 + ··· .
(13.5)
We could then plug this into the t
= 2 version of (13.2) to find that
![]() (2) = c1(l1)2
(2) = c1(l1)2 ![]() 1 + c2(l2)2
1 + c2(l2)2 ![]() 2 + ··· ,
2 + ··· ,
(13.6)
and so. In general, this then gives
![]() (t) = c1(l1)t
(t) = c1(l1)t ![]() 1 + c2(l2)t
1 + c2(l2)t ![]() 2 + ··· .
2 + ··· .
(13.7)
b) Transition/Markov
matrices
The
matrix A that appears in (13.2) has entries that are conditional probabilities,
and this has the following implications:
∑
All entries are non-negative.
∑
A0,m
+ A1,m + ··· + AN,m = 1 for all m Î {0, 1, …, N}.
(13.8)
The last line above asserts that
the probability is 1 of there being some number, either 0 or 1 or 2, or
… or N of red cells in the subsequent generation given m red cells in the
initial generation.
A square
matrix with this property is called a transition
matrix, or sometimes a Markov matrix. When A is such a matrix, the equation in
(13.2) is called a Markov process.
Although
we are interested in the eigenvalues of A, it is amusing to note that the
transpose matrix, AT, has an eigenvalue equal to 1 with the
corresponding eigenvector being proportional to the vector, ![]() , with each entry equal to 1.
Indeed, the entry in the m’th row and n’th column of AT is Anm,
this the entry of A in the n’th row and m’th column. Thus, the m’th entry of AT
, with each entry equal to 1.
Indeed, the entry in the m’th row and n’th column of AT is Anm,
this the entry of A in the n’th row and m’th column. Thus, the m’th entry of AT![]() is
is
(AT![]() )m = A0,ma0 + A1,ma1
+ ··· + AN,maN .
)m = A0,ma0 + A1,ma1
+ ··· + AN,maN .
(13.9)
If the lower line in (13.8) is
used and if each ak is 1, then each entry of AT![]() is also 1. Thus, AT
is also 1. Thus, AT![]() =
= ![]() .
.
As
a ‘cultural’ aside, what follows is the story on Anm in the example
from Part a. First, Anm = 0
of n is larger than 2m since m parent cells can spawn at most 2m
survivors. For n ≤ m, consider
that you have 2N cells of which 2m are red and you ask for the probability that
a choice of N from the 2N cells results in n red ones. This is a counting problem that is much like
those discussed in Part c of Handout 9 although more complicated. The answer here is:
Anm = ![]() when 0 ≤ n ≤ 2m.
when 0 ≤ n ≤ 2m.
(13.8)
c) Another protein
folding example
Here
is another model for protein folding. As
you may recall from previous handouts, a protein is made of segments tied end
to end as a chain. Each segment is one
of 20 amino acids. The protein is made
by the cell in a large and complex molecule called a ribosome. The segments are attached one after the other
in the ribosome and so the chain grows, link by link. Only a few segments are in the ribosome, and
the rest stick out as the protein grows.
As soon as a joint between two segments is free of the ribosome, it can
bend if it is not somehow stabilized by surrounding molecules. Suppose that the bend at a joint can be in
one of n directions as measured relative to the direction of the previously
made segment. A simplistic hypothesis
has the direction of the bend of any given segment influenced mostly by the
bend direction of the previously made segment.
To model
this, let me introduce, for k, j Î {1, …, n}, the conditional
probability, Ak,j that the a given segment has bend direction k when
the previously made segment has bend direction k´. Next, agree to label the segments along the
protein by the integers in the set {1, …, N}.
If z denotes such an integer, let pk(z) denote the
probability that the z’th segment is bent in the k’th direction relative to the
angle of the z-1’st segment. Then we
have
pk(z) = ∑1≤j≤n Akj
pj(z-1)
(13.9)
I can now introduce the vector ![]() (z) in Rn whose k’th component is pk(z), and also the
square matrix A with the components Akj. Then (13.9) is the equation
(z) in Rn whose k’th component is pk(z), and also the
square matrix A with the components Akj. Then (13.9) is the equation
![]() (z) = A
(z) = A![]() (z-1).
(z-1).
(13.10)
Note that as in the previous case,
the matrix A is a Markov matrix. This is
to say that each Akj is non-negative because they are conditional
probabilities; and
A1,j + A2,j + ··· An,j = 1
(13.11)
because the segment be at some
angle or other.
Here
is an explicit example: Take n = 3 so
that A is a 3 ´ 3 matrix. In particular,
take
A = 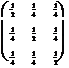 .
.
(13.12)
As it turns out, A has the
following eigenvectors:
![]() 1 =
1 = ![]()
![]() ,
, ![]() 2 =
2 = ![]()
![]() and
and ![]() 3 =
3 = ![]()
![]()
(13.13)
with corresponding eigenvalues l1 = 1, l2 = ![]() and l3 =
and l3 = ![]() . Note that the
vectors in (13.13) are mutually orthogonal and have norm 1, so they define an
orthonormal basis for R3. Thus, any given z
version of
. Note that the
vectors in (13.13) are mutually orthogonal and have norm 1, so they define an
orthonormal basis for R3. Thus, any given z
version of ![]() (z) can be written as a linear combination of the vectors in
(13.13). Doing so, we write
(z) can be written as a linear combination of the vectors in
(13.13). Doing so, we write
![]() (z) = c1(z)
(z) = c1(z) ![]() 1 + c2(z)
1 + c2(z) ![]() 2 + c3(z)
2 + c3(z) ![]() 3
3
(13.14)
where each ck(z) is
some real number. In this regard, there
is one point to make straight away, which is that c1(z) must equal ![]() when the entries of
when the entries of ![]() (z) represent probabilities that sum to 1. To explain, keep in mind that the basis in
(13.13) is an orthonormal basis, and this implies that
(z) represent probabilities that sum to 1. To explain, keep in mind that the basis in
(13.13) is an orthonormal basis, and this implies that ![]() 1·
1·![]() = c1(z).
However, since each entry of
= c1(z).
However, since each entry of ![]() 1 is equal to
1 is equal to ![]() , this dot product is
, this dot product is ![]() (p1(z) + p2(z) + p3(z)) and
so equals
(p1(z) + p2(z) + p3(z)) and
so equals ![]() when p1+p2+p3
= 1 as is the case when these coefficients are probabilities.
when p1+p2+p3
= 1 as is the case when these coefficients are probabilities.
In
any event, if you plug the expression in (13.14) into the left side of (13.10)
and use the analogous z-1 version on the right side, you will find that the
resulting equation holds if and only if the coefficients obey
c1(z) = c1(z-1) , c2(z) = ![]() c2(z-1) and
c3(z) =
c2(z-1) and
c3(z) = ![]() c3(z-1).
c3(z-1).
(13.15)
Note that the equality here
between c1(z) and c1(z-1) is heartening in as much as
both of them are supposed to equal ![]() . Anyway, continue by iterating (13.15) by
writing the z-1 versions of ck in terms of the z-2 versions, then
the latter in terms of the z-3 versions, and so on until you obtain can be
iterated now to find that
. Anyway, continue by iterating (13.15) by
writing the z-1 versions of ck in terms of the z-2 versions, then
the latter in terms of the z-3 versions, and so on until you obtain can be
iterated now to find that
c1(z) = c1(0), c2(z)
= (![]() )z c2(0) and c3(z)
= (
)z c2(0) and c3(z)
= (![]() )z c3(0).
)z c3(0).
(13.16)
Thus, we see from (13.16), we have
![]() (z) =
(z) = ![]()
![]() + (
+ (![]() )z
)z 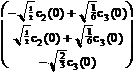 .
.
(13.17)
By the way, take note of how the probabilities for the three possible fold directions come closer and closer to being equal as z increases even if the initial z = 0 probabilities were drastically skewed to favor one or the other of the three directions. For example, suppose that
![]() (0) =
(0) = ![]() .
.
(13.18)
As a result, c1(0) = ![]() , c2(0) = -
, c2(0) = -![]() and c3(0) =
and c3(0) = ![]() . Thus, we have
. Thus, we have
![]() (z) =
(z) = ![]()
![]() + (
+ (![]() )z
)z ![]() .
.
(13.8)
Thus, by z = 4, the three
directions differ in probability by less than 1%.
Exercises
1.
Multiply the matrix A in (3.12) against the vector ![]() (z) in (13.17) and verify that the
(z) in (13.17) and verify that the
result is equal to ![]() (z+1) as defined by replacing z by z+1 in (13.17).
(z+1) as defined by replacing z by z+1 in (13.17).
2. Let A denote the 2 ´ 2
matrix ![]() . The vectors
. The vectors ![]() 1 =
1 =![]()
![]() and
and ![]() 2 =
2 =![]()
![]()
are eigenvectors of A.
a) Compute the eigenvalues of ![]() 1 and
1 and ![]() 2.
2.
b) Suppose
z Î {0, 1, …} and ![]() (z+1) = A
(z+1) = A![]() (z) for all z. Find
(z) for all z. Find ![]() (z) if
(z) if ![]() (0) =
(0) = ![]() ?
?
c) Find ![]() (z) if
(z) if ![]() (0) =
(0) = ![]() ?
?
d)
Find![]() (z) in the case that
(z) in the case that ![]() (z+1) =
(z+1) = ![]() + A
+ A![]() (z) all z and
(z) all z and ![]() (0) =
(0) = ![]() .
.
3.
Suppose that you have a model to explain your data that predicts the probability
of a
certain
measurement having any prescribed value.
Suppose that this probability function has mean 1 and standard deviation
2.
a) Give an
upper bound for the probability of a measurement being greater than 5.
b) Suppose that you average some very large
number, N, of measurements that are
taken
in unrelated, but identical versions of the same experimental set up. Write
down a
Gaussian probability function that you can use to estimate the probability that
the value of this average is greater than 5.
In particular, give a numerical estimate using this Gaussian function
for N = 100.
c) Lets say that you will agree to throw out your proposed model if it predicts that the
probability
for finding an average value that is greater than your measured average for
100 measurements is less than ![]() . If your measured
average is 1.6 for 100 experiments, should you junk your model?
. If your measured
average is 1.6 for 100 experiments, should you junk your model?
4. Suppose that you repeat the same experiment 100 times and each time record the value
of a
certain key measurement. Let {x1,
…, x100} denote the values of the measurement in N experiments. Suppose that ∑1≤k≤100
xk = 0 and ![]() ∑1≤k≤100 xk2
= 1. Suppose, in addition, that five of
the xk obey xk > 3.
The purpose of this problem is to walk you through a method for
obtaining an upper bound for the likelyhood of having five measurements that
far from the mean. In this regard,
suppose that you make the hypothesis that the spread of the measured numbers {xk}1≤k≤100
is determined by the Gaussian probability function with mean 0 and standard
deviation equal to 1.
∑1≤k≤100 xk2
= 1. Suppose, in addition, that five of
the xk obey xk > 3.
The purpose of this problem is to walk you through a method for
obtaining an upper bound for the likelyhood of having five measurements that
far from the mean. In this regard,
suppose that you make the hypothesis that the spread of the measured numbers {xk}1≤k≤100
is determined by the Gaussian probability function with mean 0 and standard
deviation equal to 1.
a) Let p denote the
probability using this Gaussian probability function for a measurement x with x
≥ 3. Explain why p ≤ ![]() 3 e-9/2 ≤ 0.014.
3 e-9/2 ≤ 0.014.
b) Use the binomial probability function to explain
why the probability of finding five or more values for k out of 100 that have xk
> 3 is equal to
∑5≥k≥100 ![]() pk (1-p)100-k.
pk (1-p)100-k.
c) Let k ≥ 5
and let z ® ƒk(z) = zk
(1-z)100-k. Show that ƒ is an
increasing function of z when z < ![]() . (Hint: Take its derivative with respect to z.)
. (Hint: Take its derivative with respect to z.)
d) Since 0.014
≤ 0.05 = ![]() , use the result from Part d to conclude that the probability
of finding 5 or more of the xk’s out of 100 with xk >
3 has probability less than
, use the result from Part d to conclude that the probability
of finding 5 or more of the xk’s out of 100 with xk >
3 has probability less than
∑5≥k≥100 ![]() (0.014)k (0.986)100-k.
(0.014)k (0.986)100-k.
e) We saw in
Equation 9.21 of Handout 9 that the terms in the preceding sum get ever smaller
as k increases. Use a calculator to show
that the k = 9 term and thus all higher k terms are smaller than 1.1´10-5. Then, use a calculator to estimate the k = 5,
6, 7 and 8 terms to prove that the sum in Part d is no greater than 0.015.