Bio/statistics Handout 5 How matrix products arise
·
Genetics:
Suppose that a given stretch of DNA
coding for cellular product is very mutable, so that there are some number, N,
of possible sequences that can appear in any given individual (this is called
‘polymorphism’) in the population. To
elaborate, a strand of DNA is a molecule that appears as a string of small,
standard molecules that are bound end to end.
Each of these standard building blocks can be one of four, labeled C, G,
A and T. The
order in which they appear along the strand determines any resulting cellular
product that the given part of the DNA molecule might produce. For example, AAGCTA may code for a different product than
GCTTAA.
As it turns out, there are stretches of DNA where the code
can be changed without damage to the individual. What with inheriting genes from both parents,
random mutations over the generations can then result in a population where the
codes on the given stretch of DNA vary from individual to individual. In this situation, the gene is called
‘polymorphic’. Suppose that there are N
different possible codes for a given stretch.
For example, if one is looking at just one particular site along a
particular DNA strand, there could be at most N = 4 possibilities at that site,
namely C, G, A or T. Looking at two
sites gives N = 4´4 = 16 possibilities.
Lets label the possible sequences by integers starting from 1 and going to
N. At any given generation t, let pj(t)
denote the frequencey of the j’th
sequence appearing in the population at that generation. Given that the j’th
sequence appears in an individual, there is some probability, P(i|j), that the i’th sequence appears in their heir. Here, let us assume that the population is
constant and that these P(i|j)
are independent of the generation number.
(This is all very simplistic except in very special circumstances.) In this situation, we can say the
following: The probability of any given
sequence, say i, appearing in generation t+1 can be
written as a sum:
pi(t+1)
= P(i|1) p1(t) + P(i|2) p2(t) + · · · + P(i|N) pN(t)
(5.1)
This is to say that the probability of sequence i appearing in an heir is equal to the probability that sequence 1 appears in the parent times the probability of a mutation that changes sequence 1 to sequence i, plus the probability that sequence 2 appears in the parent times the probability of a mutation that changes sequence 2 to sequence i, etc.
Now,we can write this using our
matrix formalism by thinking of the numbers {pj(t)}1≤j≤N
as defining a column vector, ![]() (t), in RN, and likewise the numbers {pi(t+1)}1≤i≤N
as defining a second column vector,
(t), in RN, and likewise the numbers {pi(t+1)}1≤i≤N
as defining a second column vector, ![]() (t+1) in RN. If I introduce the N´N matrix A whose entry in the j’th
column and i’th row is P(i|j), then (5.1) says in very cryptic shorthand:
(t+1) in RN. If I introduce the N´N matrix A whose entry in the j’th
column and i’th row is P(i|j), then (5.1) says in very cryptic shorthand:
![]() (t+1) = A
(t+1) = A![]() (t) .
(t) .
(5.2)
Now, we can sample the population at time t = T = now, and
thus determine ![]() (T), or at least the proxy that takes pi(T) to be
the percent of people in the population today that have sequence i. One very
interesting question is to determine
(T), or at least the proxy that takes pi(T) to be
the percent of people in the population today that have sequence i. One very
interesting question is to determine ![]() (T´) at some point far in the past, thus T´ << T. For example, if we find
(T´) at some point far in the past, thus T´ << T. For example, if we find ![]() (T´) such that all pi(T´)
are zero but a very few, this then indicates that the population at time T´ was
extremely homogeneous, and thus presumably very small.
(T´) such that all pi(T´)
are zero but a very few, this then indicates that the population at time T´ was
extremely homogeneous, and thus presumably very small.
To determine ![]() (T´), we use (5.2) in an iterated
form:
(T´), we use (5.2) in an iterated
form:
![]() (T) = A
(T) = A![]() (T-1) = AA
(T-1) = AA![]() (T-2)) = AAA
(T-2)) = AAA![]() (T-3) = ··· =
A···A
(T-3) = ··· =
A···A![]() (T´) ,
(T´) ,
(5.3)
where the final term has T-T´ copies of A multiplying one after the other.
On a similar vein, we can use (5.2) to predict the distribution of the sequences in the population at any time T´ > T by iterating it to read
![]() (T´) = A···A
(T´) = A···A![]() (T)
(T)
(5.4)
Here, the multiplication is by T´-T
successive copies of the matrix A.
By the way, here is a bit of notation: The sequence {![]() (t)}t=0,1,… of
vectors of probabilities is an example of a Markov
chain. In general, a Markov chain is
a sequence of probabilities, {P(0), P(1), P(2), …, }
where the N’th probability P(N) depends only on the
probabilities with numbers that are less than N.
(t)}t=0,1,… of
vectors of probabilities is an example of a Markov
chain. In general, a Markov chain is
a sequence of probabilities, {P(0), P(1), P(2), …, }
where the N’th probability P(N) depends only on the
probabilities with numbers that are less than N.
·
How bacteria find food: If you put certain
sorts of bacteria on one side of a petri dish and put
some sugar some distance away on the other, the bacteria will migrate towards
the sugar. Apparently, the sugar
diffuses in the petri dish, so that there is slight concentrations everywhere, with most of the
concentration in the original spot. The
bacteria are sensitive to the different levels at their front and rear ends and
tend to move in the direction of the greater level. At the expense of borrowing from Math 21a,
the bacteria sense the direction of the gradient
of the sugar concentration. Even so,
their movement from low to high concentration has a certain
randomness to it; at any given step there is some probability that they will
move in the wrong direction.
What follows is a simplistic model for this: Imagine bacteria moving in steps along the
x-axis along the segment that stretches from x = 1 to x = N. Here, N is some large integer. Suppose the sugar placed initially where x =
N, and the bacteria is placed initially at x = 1. Our model also supposes that the bacteria
moves one unit per second, with probability q Î (0, 1) of moving to the
right at any given step, and probability 1-q of moving to the left unless it is
at the end of the interval. If it is at
the x = 1 end, it moves to the right with probability q and stays put with
probability 1-q. If it is at the x = N
end, it stays put with probability q and moves to the left with probability
1-q. In our model, q is independent of
position, x Î {1, . . . , N}. Such would be roughly the case in the real petri dish where the sugar concentration gradient is
relatively constant. We should take q
> ![]() in the case that the
bacteria is attracted to the sugar.
in the case that the
bacteria is attracted to the sugar.
For each time step t, and j Î {1, .
. . , N}, let pj(t) denote the
probability that the bacteria is at position j at time t. For example, p1(0)
= 1 and pj(0) = 0 for j > 1. Our model then says that the probability of
finding the bacteria at position j ≠ 1 of N at time step t > 0 is
equal to the probability that it is at position j-1 at time t-1 times the probability
that it moves one step to the right, plus the probability that it is at
position j+1 at time t-1 and moves one step to the left. Thus,
pj(t+1) = q pj-1(t)
+ (1-q) pj+1(t) when 2
≤ j ≤ N-1
(5.5)
By the same token,
p1(t+1)
= (1-q) p1(t) + (1-q) p2(t) and pN(t+1)
= q pN-1(t) + q pN(t) .
(5.6)
Let us introduce the vector ![]() (t) in RN whose j’th
component is pj(t). Then (5.5) and
(5.6) assert that
(t) in RN whose j’th
component is pj(t). Then (5.5) and
(5.6) assert that ![]() (t+1) is obtained from
(t+1) is obtained from ![]() (t) by the action of a linear transformation:
(t) by the action of a linear transformation: ![]() (t+1) = A
(t+1) = A![]() (t), where A is the N´N
matrix whose only non-zero entries are:
(t), where A is the N´N
matrix whose only non-zero entries are:
A11 = 1-q,
Ajj-1 = q for 2 ≤ j ≤ N, Ajj+1 = 1-q for 1
≤ j ≤ N-1, and ANN
= q
(5.7)
For example, in the case that N = 4, the
matrix A is
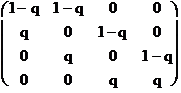
(5.8)
In any case, we can iterate the equation ![]() (t+1) = A
(t+1) = A![]() (t) to find that
(t) to find that
![]() (t) = A···A
(t) = A···A![]() (0)
(0)
(5.9)
where
A···A signifies t copies of A multiplied one after the other. By the way, a common
shorthand for some n copies of any given matrix, M, successively multiplying
one after the other is Mn.
·
Growth of nerves in a developing embryo: Here is a very
topical question in the study of development:
Nerve cells connect muscles and organs to the brain. When an organism develops, how do its nerves
know where to connect? A given nerve
cell stretches out extremely long and thin ‘appendages’ called axons. Any given axon may end abutting a muscle
cell, or another nerve cell in a chain of nerves that ends in the brain. How do the axons ‘know’ where they are
supposed to attach? A simple model proposes that the tip of the growing axon in an embryo
is guided by chemical gradients in much the same fashion as the bacteria in the
previous discussion is guided to the sugar.
·
Enzyme dynamics: An enzyme is a
protein molecule that facilitates a chemical reaction that would take a long
time to occur otherwise. Most biological
reactions are facilitated by enzymes.
One typical mode of operation is for the enzyme to facilitate a reaction
between molecules of type a and molecules of type b to
produce a new molecule of type g. It can do this if it simulaneously
attracts the a and b molecules. In doing
so, the enzyme holds the two sorts near each other for enough time that they
can bind together to form the more complicated g molecule. If this g molecule does noot have a strong attraction to the enzyme, it will break
away and so free the enzyme to attract another a and another b to
make more g.
Now, it is often the case that the enzyme only works efficently if it is folded in the right
conformation. Folded in the wrong way,
the parts of the molecule that attract either a or b
molecules might find themselves covered by parts that are indifferent to a or b. Due to random collisions and other effects,
any given molecule, and thus our enzyme, will change its fold
configuration. Suppose that there is
some probability, q, that the enzyme is folded correctly to attract a and b in
any given unit of time, and thus probability (1-q) that it is not. Suppose in addition, that when folded
correctly, the enzyme makes 2 units of g per unit of time. Meanwhile, suppose that the freely diffusing g falls
apart or is degraded by other enzymes at a rate of 1 unit per unit time.
For any integer j, let pj(t) denote the
probability of finding level j of g after some t units of time.
Then the scenario just outlined finds
pj(t+1) = q pj-1(t)
+ (1-q) pj+1(t) when j
≥ 1
and p0(t+1) =
(1-q)(p1(t) + p0(t))
(5.10)
as long as a and b are well supplied.
Exercises:
Exercises 1-3 concern the example
above where the position of the bacteria at any given time t is labeled by an
integer, x(t), in the set {1, . . . , N}.
1.
Suppose
T is a positive integer.
a) What is the sample space for the collection of
positions, {x(0), x(1), . . . , x(T)}, of the bacteria
at the times t = 0, 1, …, T.
b) Fix some j Î {1, …,
N} and t Î {0, 1, …, T-1} and let A denote the event that x(t+1) =
j. For each i Î {1, …, N}, let Aj denote
the event that x(t) = i. Explain why the equation in (5.5) has the
form P(A) = P(A|A1)P(A1) + ···
+ P(A|An)P(AN).
2.
Recall
that events A and B are said to be independent when P(AÇB) =
P(A)P(B).
a) Explain why this definition is equivalent to
the assertion that the conditional probability of A
happening given B is equal to the probability of A happening.
b) Suppose that k Î {2, . . . , N-1} and that t ≥ 2. Derive a relation between the respective
probabilities that x(t) = k-1 and x(t) = k+1 so as to
insure that the event that x(t+1) = k is independent of the event that x(t) =
k-1.
3.
Suppose
now that q+ and q- are non-negative numbers whose sum is
less than 1. Change the bacteria model
so that if the bacteria is at position k Î {2, . . . , N-1} at time
t, then it moves to position k+1 with probability q+, to position
k-1 with probability q- and stays put with probability 1-q+-q-. If the bacteria is at position 1 at time t,
then it stays put with probability 1-q+ and moves to position 2 with
probability q+. If the
bacteria is at position N at time t, then in stays put with probability 1-q-
and moves to position N-1 with probability q-
a) Write
down the analog of (5.5) for this model.
b) Write
down the analog of the matrix in (5.8) for the N = 4 case.
c) Write down the
probability that the bacteria is at position N at t = N-1 given that the
bacteria starts at position 1 at time 0.
4.
Write
down a version of (5.10) that would hold in the case that the enzyme when
folded correctly produces some L = 1, or L > 2 units of g per
unit time.